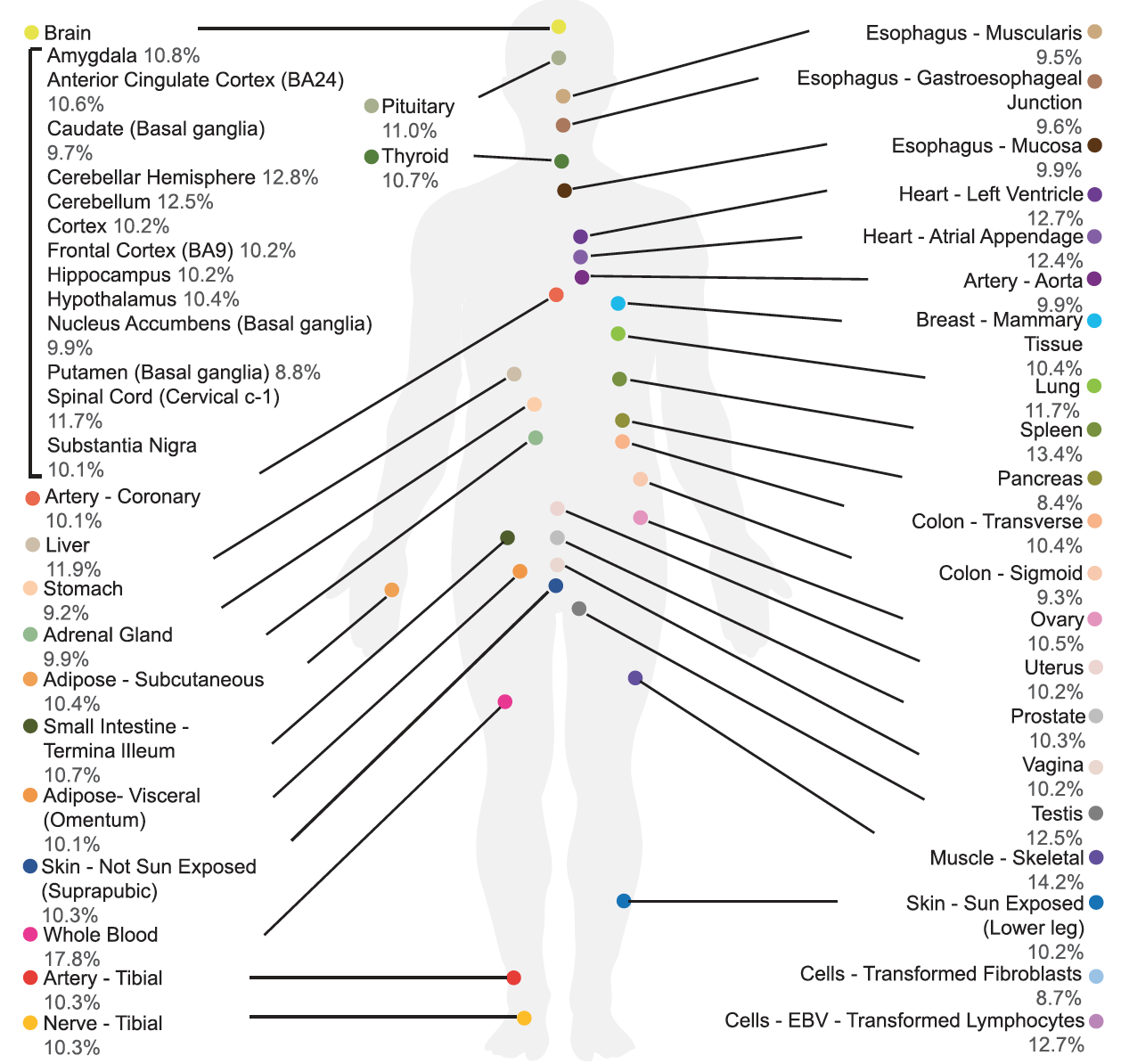
We are a hybrid lab of computational and experimental RNA biology. The overarching goals of our research are to understand how the transcriptome and epitranscriptome are controlled by the intricate network of genetic factors, RNA elements, and RNA-binding proteins, and how such regulation contributes to neurological diseases and cancer.
RNA regulation linking genetics and disease
Genetic variants can contribute to human diseases in many ways. Most previous studies focused on their impact on protein-coding sequences or transcriptional regulation. In recent years, we and others have revealed numerous genetic variants that alter post-transcriptional regulation. Our lab is focusing on the genetic regulation of splicing and RNA stability.
Key papers:
scAllele: a versatile tool for the detection and analysis of variants in scRNA-seq
Quinones-Valdez G, Fu T, Chan TW, Xiao X.
Science Advances. Vol 8, Issue 35, 2 September 2022.
Allele-specific alternative splicing and its functional genetic variants in human tissues
Amoah K, Hsiao YHE, Bahn JH, Sun Y, Burghard C, Tan BX, Yang EW, Xiao X.
Genome Research, Published in Advance January 15, 2021, doi:10.1101/gr.265637.1
The epitranscriptome and its regulation
The epitranscriptome refer to the collection of post-transcriptional RNA modifications. Novel discoveries and technological advancements over the past decade have catalyzed this young field of epitranscriptomics. Our lab is currently focusing on RNA editing, a type of RNA modification that alters A to I.
Key papers:
Single-cell analysis in lung adenocarcinoma implicates RNA editing in cancer innate immunity and patient prognosis. Chan TW, Dodson JP, Arbet J, Boutros PC, Xiao X.
Cancer Research. 2022 Nov 30;CAN-22-1062.
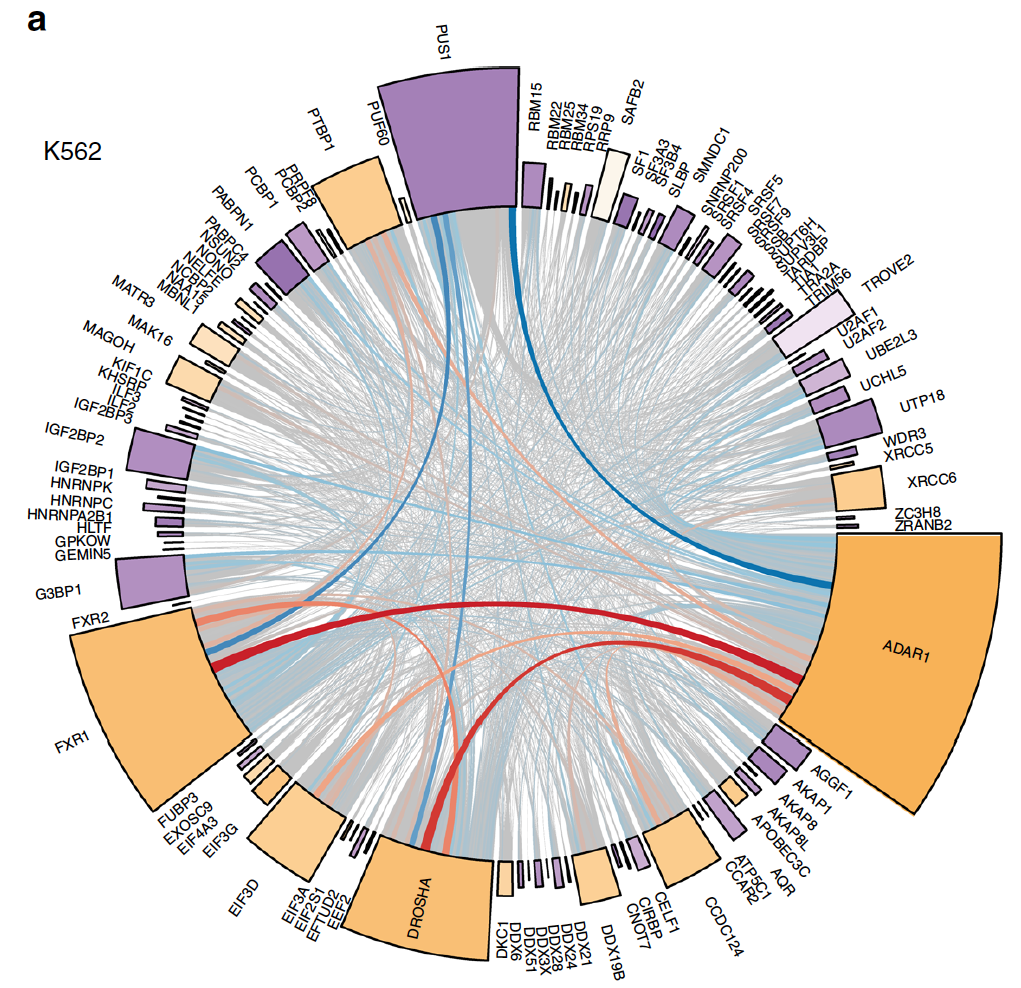
Regulation of RNA editing by RNA-binding proteins in human cells.
Quinones-Valdez G, Tran SS, Jun HI, Bahn JH, Yang EW, Zhan L, Brümmer A, Wei X, Van Nostrand EL, Pratt GA, Yeo GW, Graveley BR, Xiao X.
Commun Biol. 2019;2:19. doi: 10.1038/s42003-018-0271-8. eCollection 2019. PubMed PMID: 30652130.